Winning solution of the EUOS25 Challenge (Transmittance Category)
In the EUOS25 Challenge[1] hosted by EU-OPENSCREEN and SLAS, our team yumiz achieved first place in the blind test of the Transmittance category (we placed 6th in the Fluorescence category). As the winning team, we will be awarded a prize of €1,000 at the SLAS2026 International Conference & Exhibition in Boston. Additionally, the winning team, as well as other teams whose models achieved an AUC not significantly different from that of the winning solution, will be invited to publish their studies or protocols in the SLAS journal. In this blog, we would like to walk through our solution and share the experiences and lessons we gained during the competition.
Competition Overview
The EUOS25 Challenge is a data science competition (similar to Kaggle) for predicting compound transmittance (absorption) and fluorescence. A similar competition was held two years ago (https://www.kaggle.com/competitions/euos-slas/).
The competition uses tens of thousands of training samples for binary classification tasks: the Transmittance task is to predict whether compounds absorb light at specific wavelengths, and the Fluorescence task is to predict whether compounds fluoresce above predefined thresholds. Rankings are determined separately for the two categories, each with two subtasks; the average ROC AUC across both subtasks is used for ranking.
- Transmittance subtask 1a: predict whether compounds have transmittance ≤70% at 340 nm
- Transmittance subtask 1b: predict whether compounds have average transmittance ≤70% in the 450–679 nm range
- Fluorescence subtask 2a: predict compounds whose fluorescence intensity exceeds the threshold at 340 nm excitation and 450 nm emission
- Fluorescence subtask 2b: predict compounds whose fluorescence intensity exceeds the threshold for at least one of excitation 480 nm / emission 540 nm, excitation 525 nm / emission 598 nm, or excitation 560 nm / emission 610 nm
Subtasks 1b and 2b are known to have severe label imbalance (only about 1.5% and 0.23% positives for transmittance and fluorescence, respectively), so building models that perform well on these subtasks was one of the main challenges of the competition.
The competition ran from October 2025 to January 2026.
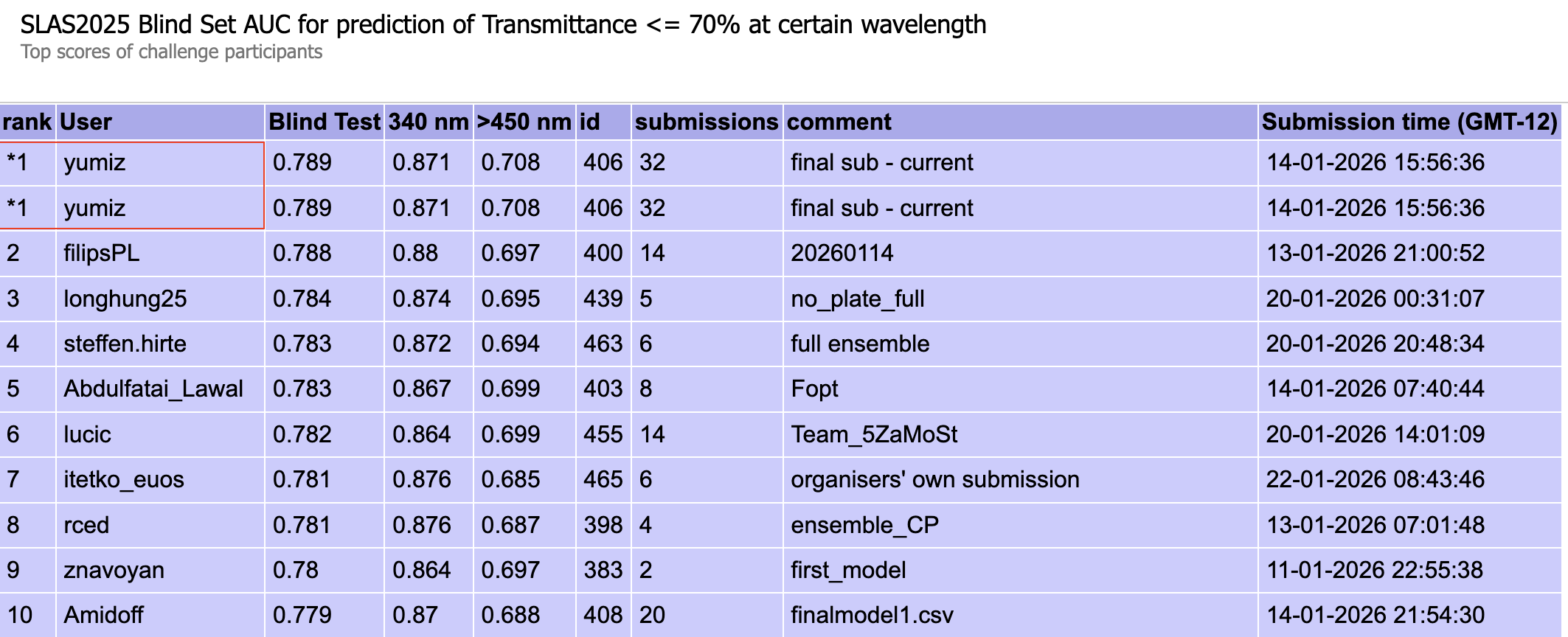
Figure 1: Transmittance blind test results (ROC AUC at 340 nm / 450–679 nm)
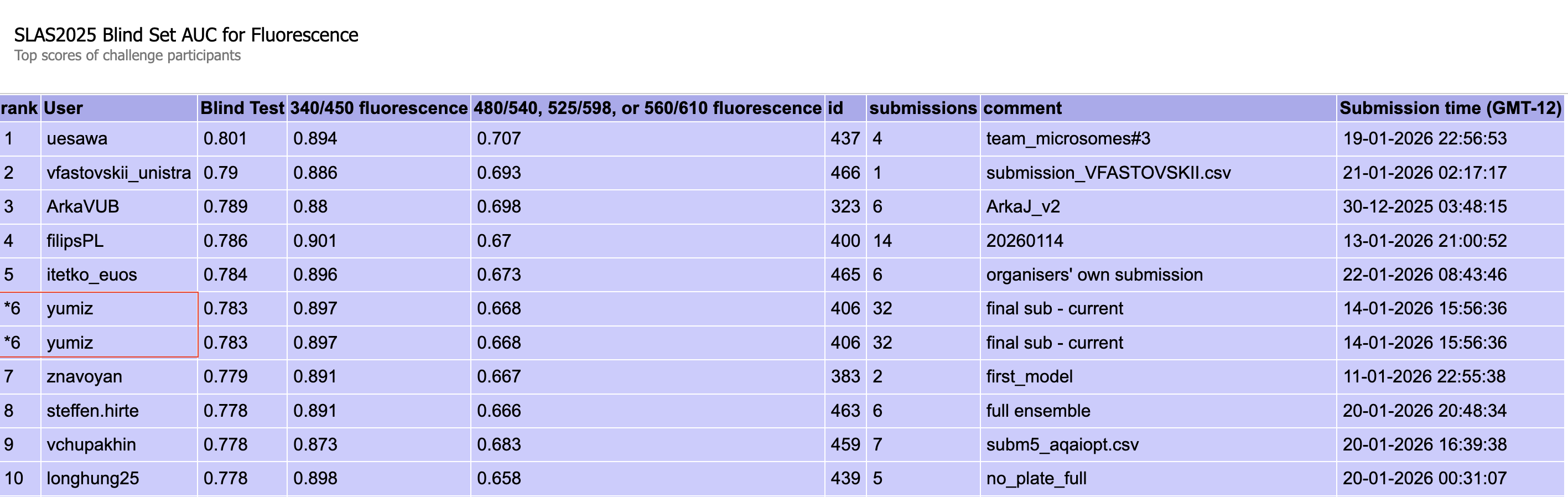
Figure 2: Fluorescence blind test results (340/450 nm and others, ROC AUC)
Solution
Here, we would like to briefly introduces our team’s solution. Multimodal Molecular Property Prediction with 1D/2D/3D Features and Weighted Ensemble. The code and implementation details are available at ohuelab/euos25-solution.
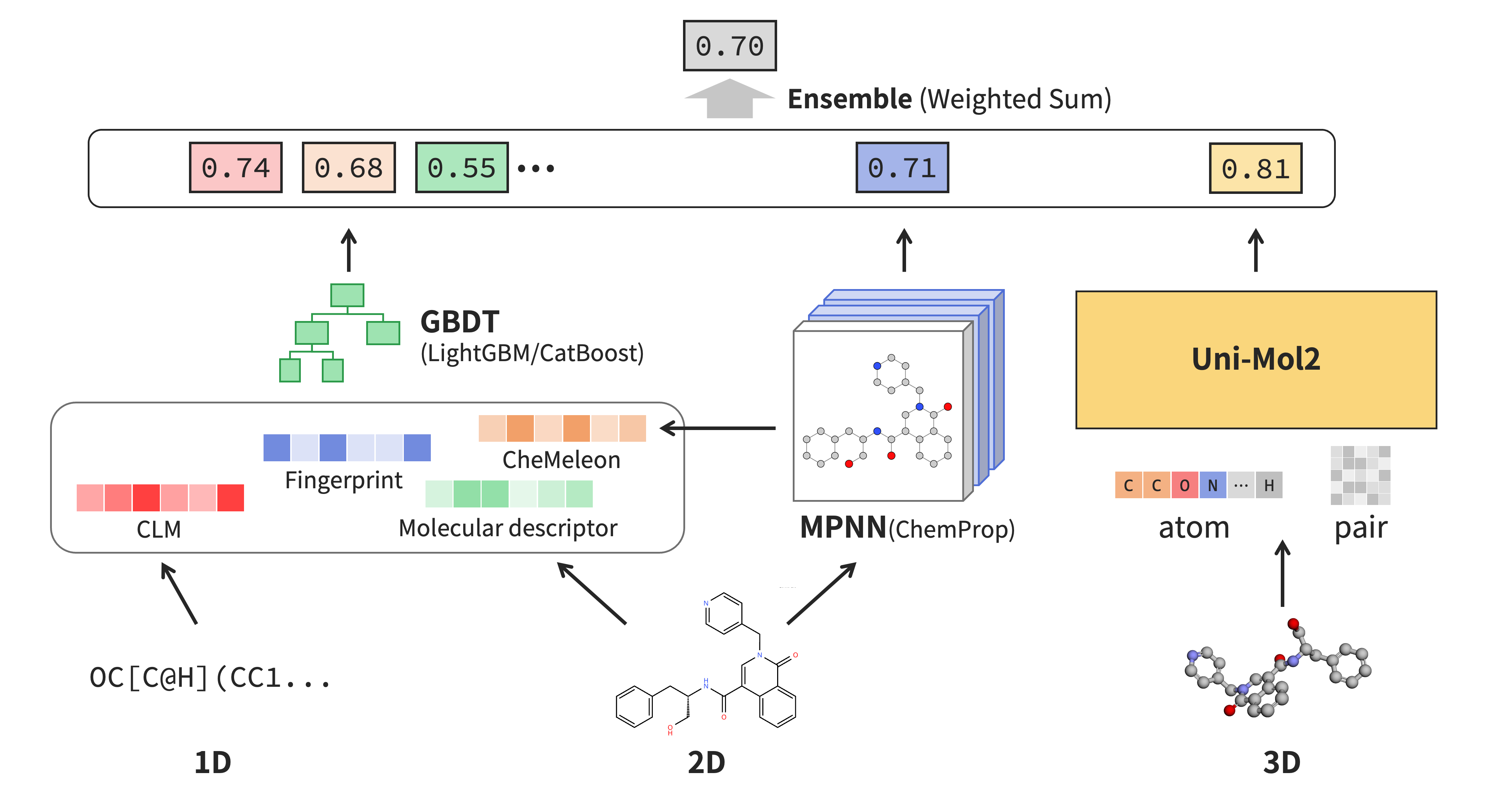
Figure 3: Overview of Team yumiz’s winning solution
In short, our solution was a weighted ensemble of multiple prediction models integrating multimodal 1D, 2D, and 3D molecular information with various model architectures.
First, we trained prediction models using gradient boosting decision trees (GBDTs) such as LightGBM and CatBoost with the following features, individually and in various combinations, and kept those with strong CV performance:
- Latent representations from a chemical language model (ChemBERTa[2])
- Molecular descriptors calculated with RDKit and Mordred
- Molecular fingerprints using ECFP4
- Latent representations from CheMeleon[3] (a ChemProp-based MPNN model pre-trained on Mordred descriptors)
We also built prediction models using ChemProp[4] and CheMeleon. Furthermore, we constructed prediction models utilizing 3D structures using Uni-Mol2[5] (84M, 310M). In order to efficiently combine multiple predictions from various models, we employed ensemble techniques with a weighted sum tuned using CV scores to select the promising combination with good accuracy. Then, we submitted the result to the system.
In summary, we used multiple-level of molecular representations including 1D information (SMILES), 2D information (structure), and 3D information (3D conformations) of molecules.
Besides the core model, we also implemented several key supporting and essential preprocessing, training, and optimization techniques:
- Basic SMILES preprocessing such as salt removal
- Considering using regression models for transmittance tasks
- Addressing imbalanced data using Focal Loss
- Multi-task learning with ChemProp
- 5-fold cross-validation with Murcko scaffolds data splitting
- Hyperparameter tuning for GBDT models using nested cross validation + Optuna
Results
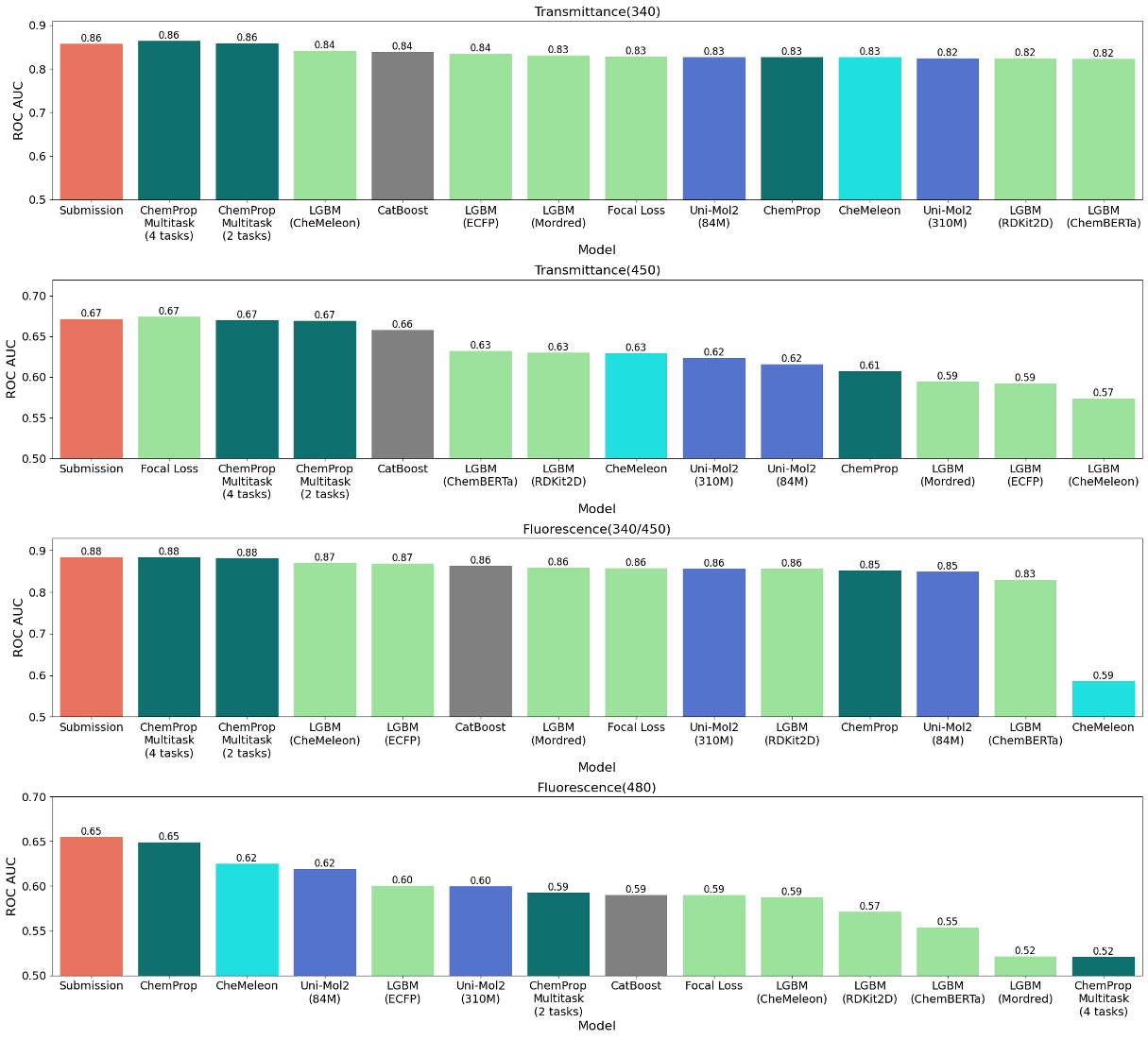
Figure 4: Performance of each method on the leaderboard for each task
The figure compares the prediction performance of multiple individual models with that of the final submitted model, using the leaderboard test set after the competition (not all individual models were used in the ensemble). While no consistent trend was observed across all tasks, the final submitted model consistently performed well. We believe that combining diverse information sources through ensembling produced a robust model that generalizes well.
Table 1 shows the public and private leaderboard scores. Private scores were higher than the public scores for every task. Although many teams ranked above us on the public leaderboard, they were likely overfitting to the public leaderboard. We had considered scaffold-based cross-validation a standard way to maintain generalization, but we were somewhat surprised by how much the rankings changed from public to private.
Table 1: Public / Private leaderboard scores
| Task | Public | Private |
|---|---|---|
| Transmittance subtask 1a | 0.857 | 0.871 |
| Transmittance subtask 1b | 0.670 | 0.708 |
| Fluorescence subtask 2a | 0.883 | 0.897 |
| Fluorescence subtask 2b | 0.655 | 0.668 |
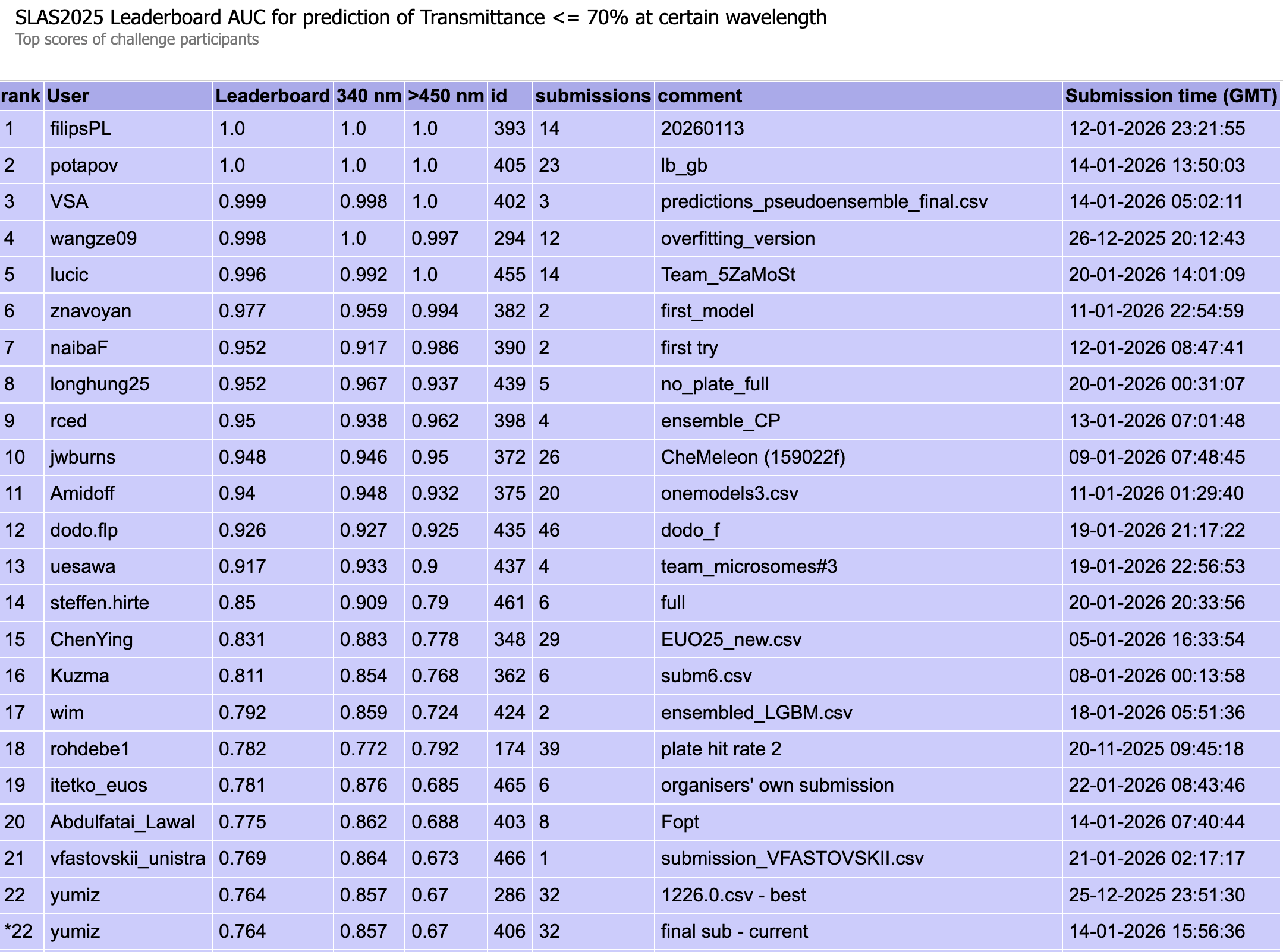
Figure 5: Public leaderboard ranking for the Transmittance task
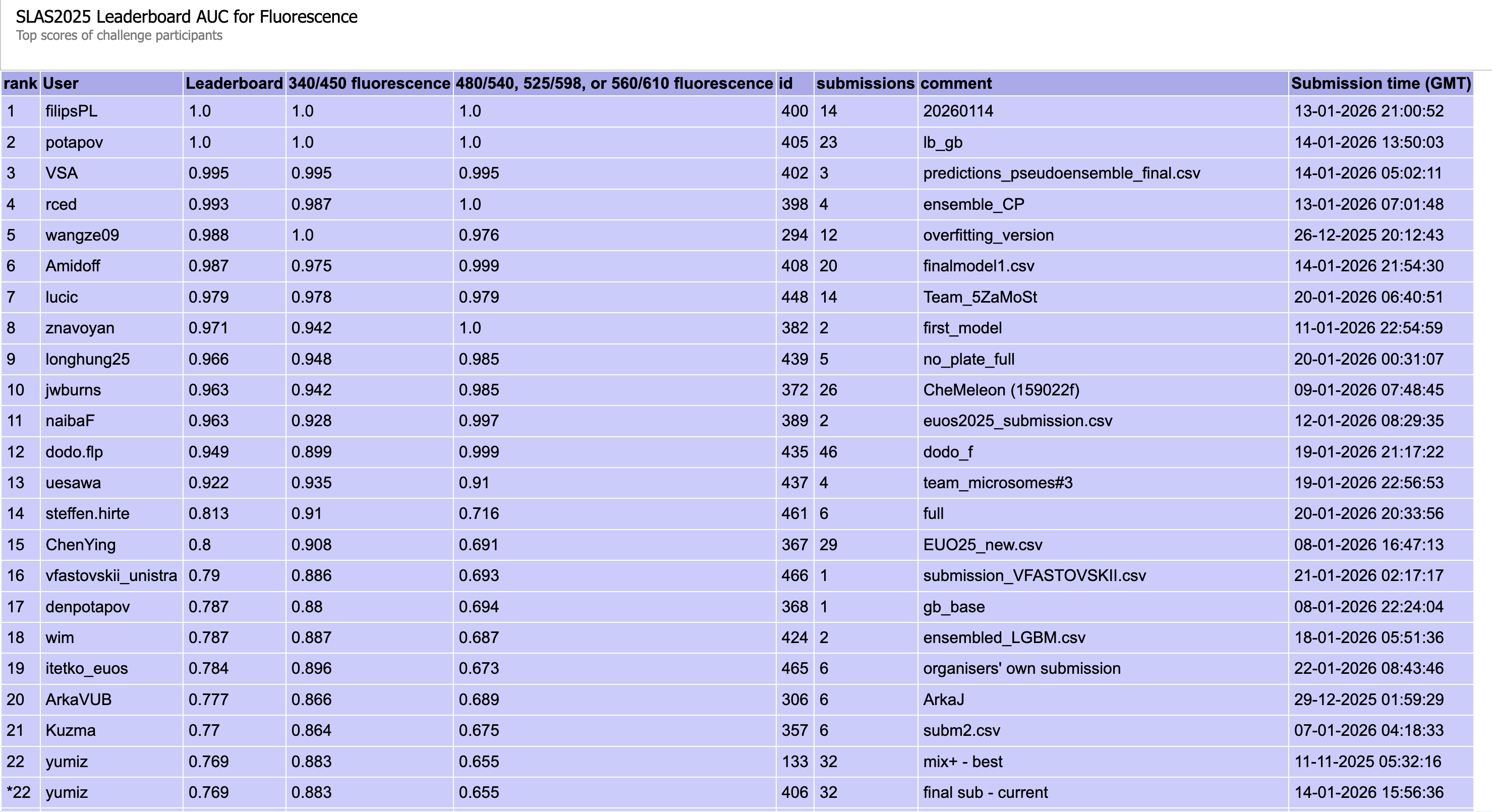
Figure 6: Public leaderboard ranking for the Fluorescence task
About Well/Plate Bias
As a side note, during the competition, our team also observed issues related to well and plate bias, which were later recognized by the organizers. From previous competitions, we have learned that such biases can arise during data splitting, and that using indices or well/plate identifiers may affect model performance. The organizers later noted that some teams reported using well/plate information to correct for distribution biases in the training data. However, although using this information for prediction was prohibited since only structural information was allowed, our own experiments showed that it was not very effective.
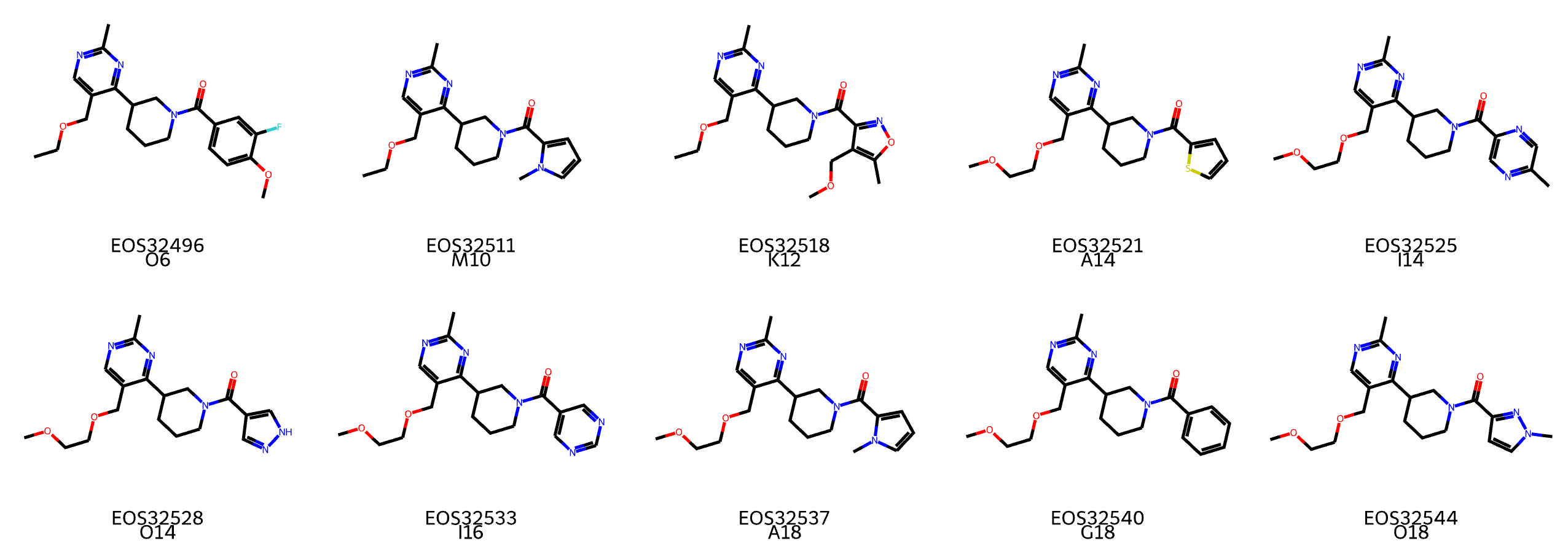
Figure 7: Very similar positive examples for 450 nm transmittance labels in plate C1096
For example, as shown in the figure, plate C1096 contains many very similar molecules that are positive for the 450 nm transmittance label. In other words, we believe the correction didn’t work because it was unclear whether the label bias in wells/plates was due to similar molecules being concentrated in the same plate, or because there were issues with those specific wells or plates.
Team Composition
Team yumiz consists of three students from Institute of Science Tokyo (Ohue Laboratory), including myself, Apakorn, and Koh. Initially, each of us participated individually, then we formed a team by contacting the organizers. We all built our own prediction model using multiple features with some ensemble methods. However, when we do ensembling, our combined three models didn’t improve performance. Thus, we decided to use my best emsembled model as the final submission. Nevertheless, forming the team was valuable, as the discussions and shared insights helped us better assess the problem and recognize that there was limited scope for further improvement.
Conclusion
In conclusion, from this competition, no single model excels across all tasks. Therefore, an ensemble approach proved to be effective. By applying weighted ensembling techniques, we were able to improve the overall robustness and stability of the model. Lastly, we’re pleased to have taken first place! The competition served as a valuable learning experience and provided useful insights for our future work. We would like to thank the competition organizers for hosting a challenging and well-structured competition. We also appreciate the efforts of the community for sharing ideas and discussions throughout the competition. We look forward to future opportunities and competitions in this space.
Acknowledgments
We thank our teammate Apakorn Kengkanna for proofreading the English in this article.
References
- [1] EUOS25 Challenge | Online Chemical Modeling Environment
- [2] Chithrananda, Seyone, et al. "ChemBERTa: Large-scale self-supervised pretraining for molecular property prediction." arXiv [cs.LG], 19 Oct. 2020
- [3] Burns, Jackson, et al. "Descriptor-Based Foundation Models for Molecular Property Prediction." arXiv [Cs.LG], 18 June 2025
- [4] Heid, Esther, et al. "Chemprop: A Machine Learning Package for Chemical Property Prediction." Journal of Chemical Information and Modeling, vol. 64, no. 1, Jan. 2024, pp. 9–17.
- [5] Ji, Xiaohong, et al. "Exploring Molecular Pretraining Model at Scale." The Thirty-Eighth Annual Conference on Neural Information Processing Systems, 2024